Gibson Assembly with CodonCode Aligner
Visualize and plan your Gibson Assembly experiment in CodonCode Aligner:
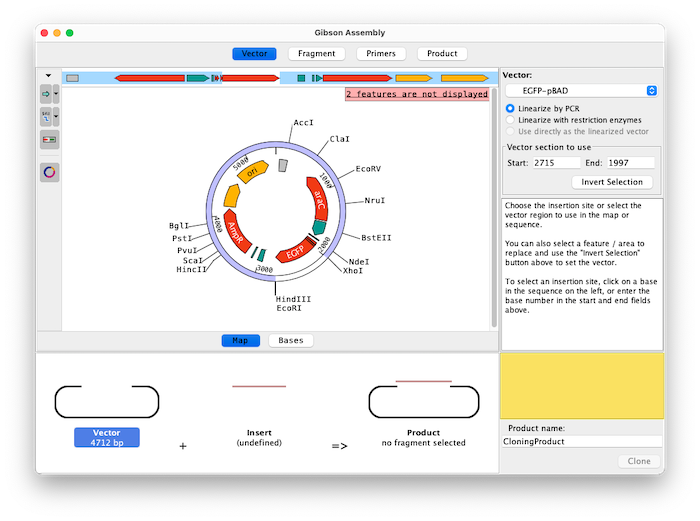
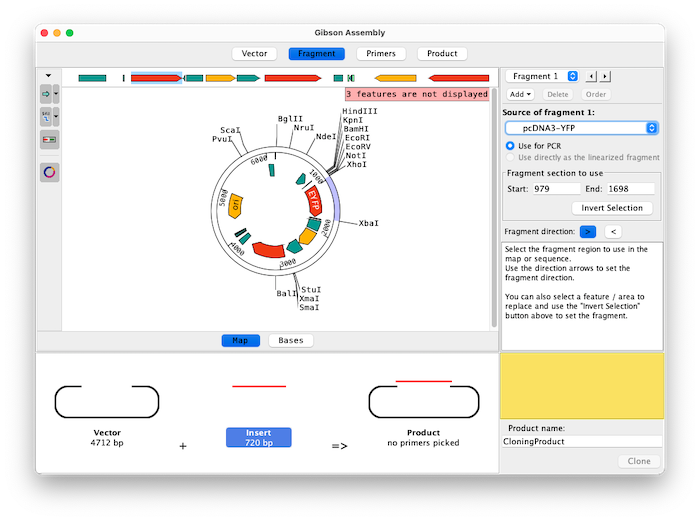
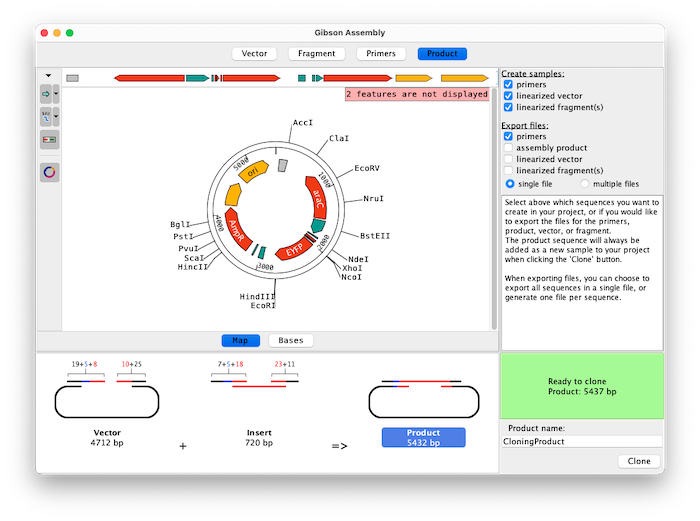
Assemble up to 15 fragments with Aligner's Gibson Assembly feature. Easily select fragments to amplify by clicking on features, selecting regions between enzymes, inverting exisiting selections, typing in a specfic range of bases, and flipping fragment directions.
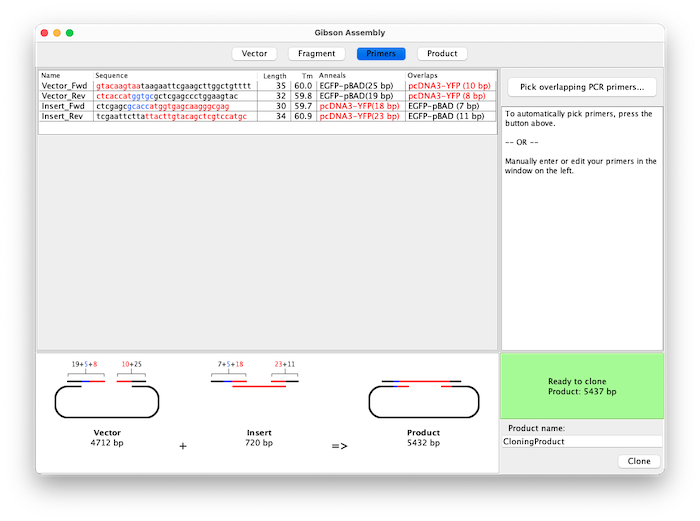
Primers are generated automatically based on your chosen overlap and Tm preferences. A table displays all primers with their information and different colors highlight the different primer sections. You can also edit primers manually - for example to add spacers or adjust the reading frame.
Switch between map and base view during the cloning product to get a great overview of all features as well as a detailed view of the bases for vector, fragments, and product.
Choose which sequences of the experiment are added to your project and export any combination of sequences and primers directly while planning your cloning experiment.
To learn more about Gibson Assembly in CodonCode Aligner, look at the Gibson Assembly tutorial.